 
Pairing Single cell TCR sequencing and RNA-seq using emulsion-based protocols. Libraries are then sequenced and TCR sequences can be extracted from all transcriptome using dedicated bioinformatics algorithms (TraCer, TraPes, VDJ Puzzle). In the library preparation step, full-length cDNA is “tagmented” using Transposase and Tag sequences inserted by transposase are then used to amplify cDNA and to insert barcoded sequencing adaptors (AD1 and AD2). This sequence, shared with the dT primer used in the RT reaction is then used to amplify cDNA before library preparation. Through a template-switch mechanism a universal sequence is added at the 5′ end of the transcript. (C) Single cells sorted in plates or captured using microfluidic devices are lysed, and total mRNA is reverse transcribed with an oligo dT primed reaction. TCR reconstruction from single cell “full length” RNA sequencing (RNA-seq) data. Fused molecules are pooled breaking the emulsion and further enriched by a nested amplification and sequenced. Alpha and beta primers contain overlapping sequences at their 5′ends that enable the synthesis of TCR alpha and beta fusion sequences through an overlap-extension mechanism. cDNA is successively amplified using a pool of forward primers designed on all the alpha and beta segments and reverse primers designed on the constant region. 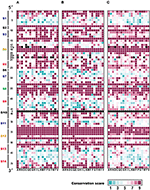
In each droplet, TCR alpha and beta transcripts of a single cell are specifically reverse transcribed with RT primers designed on the constant region of alpha and beta chain. 
In (B) cells are captured in water-in-oil droplets using microfluidic emulsion-based devices along with specific RT and PCR reagents. Barcoded adaptors then are added by PCR enabling pooling and sequencing by Next-Generation Sequencing. In (A) single cell TCR transcripts are enriched by a multiplex PCR performed after RT reaction using a pool of forward primers spanning all the annotated productive V alpha and V beta fragments and reverse primers designed on the constant region of alpha and beta chains. Overview of the available single cell T cell receptor (TCR) sequencing approaches. RNA sequencing T cell receptor repertoire bioinformatics immune system single cell analysis. We also discuss novel approaches that through the integration of TCR tracking and mRNA single cell sequencing offer a valuable tool to associate antigen specificity to transcriptional dynamics and to understand the molecular mechanisms of T cell plasticity. Single cell based approaches brought the analysis to a higher level of complexity and now provide the opportunity to sequence paired alpha and beta chains. In this review, we provide an overview of the strategies used to investigate the TCR repertoire from the pioneering techniques based on the V segments identification to the revolution introduced by Next-Generation Sequencing that allows for high-throughput sequencing of alpha and beta chains. TCRs are cell specific and represent a sort of "molecular tag" of T cells and have been widely studied to monitor the dynamics of T cells in terms of clonality and diversity in several contexts including lymphoid malignancies, infectious diseases, autoimmune diseases, and tumor immunology. This ability is achieved during thymic development through a complex molecular mechanism based on somatic recombination that leads to the expression of a very heterogeneous population of surface antigen receptors, the T Cell Receptors (TCRs). The peculiarity of T cell is their ability to recognize an infinite range of self and foreign antigens. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
